Factorial DOE: oil-company experiment#
Industrial designed experiment from an oil company. Four blend components are varied across 19 runs to improve volumetric heat capacity. Three of the components (A, B, D) are continuous and were varied in coded units; the fourth (C) is a binary factor: present or absent. The aim is to identify which factors and interactions actually drive the response so that future experimentation can focus there.
Data. oil-company-doe.csv from openmv.net. 19 rows, five columns: A, B, C, D, y.
What we do. Run the package’s analyze_experiment with an interactions model, read the coefficient table and ANOVA, build a Pareto plot of standardised effects, and check the residual diagnostics.
Adapted from the Design and analysis of experiments chapter of the Process Improvement using Data book (CC BY-SA 4.0).
[1]:
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from process_improve.experiments.analysis import analyze_experiment
Load and inspect#
[2]:
url = "https://openmv.net/file/oil-company-doe.csv"
doe = pd.read_csv(url, index_col=0)
if doe.shape[1] < 5:
doe = pd.read_csv(url)
doe.columns = [c.strip() for c in doe.columns]
print(f"Shape: {doe.shape}")
print(f"Columns: {list(doe.columns)}")
doe.head()
Shape: (19, 5)
Columns: ['A', 'B', 'C', 'D', 'y']
[2]:
| A | B | C | D | y | |
|---|---|---|---|---|---|
| 0 | 4.38 | 1.60 | No | 2.18 | 28.03 |
| 1 | 6.20 | 2.26 | Yes | 2.18 | 37.25 |
| 2 | 6.20 | 2.26 | Yes | 2.18 | 37.22 |
| 3 | 4.38 | 2.26 | Yes | 3.08 | 37.06 |
| 4 | 6.20 | 1.60 | Yes | 3.08 | 39.23 |
[3]:
doe.describe(include="all")
[3]:
| A | B | C | D | y | |
|---|---|---|---|---|---|
| count | 19.000000 | 19.000000 | 19 | 19.000000 | 19.000000 |
| unique | NaN | NaN | 2 | NaN | NaN |
| top | NaN | NaN | No | NaN | NaN |
| freq | NaN | NaN | 11 | NaN | NaN |
| mean | 5.562105 | 1.970526 | NaN | 2.685263 | 33.347895 |
| std | 0.795792 | 0.311582 | NaN | 0.424884 | 4.037329 |
| min | 4.380000 | 1.600000 | NaN | 2.180000 | 28.030000 |
| 25% | 4.680000 | 1.600000 | NaN | 2.180000 | 29.565000 |
| 50% | 6.200000 | 2.040000 | NaN | 2.780000 | 31.710000 |
| 75% | 6.200000 | 2.260000 | NaN | 3.080000 | 37.140000 |
| max | 6.200000 | 2.260000 | NaN | 3.080000 | 39.230000 |
Fit the interactions model#
analyze_experiment accepts a design DataFrame and the name of the response column, builds a formula, fits it via statsmodels, and returns a dictionary keyed by the requested analysis types. With model="interactions" we ask for main effects plus all two-factor interactions; that is 4 + 6 = 10 terms, which fits comfortably in 19 runs.
[4]:
result = analyze_experiment(
doe,
response_column="y",
model="interactions",
analysis_type=[
"coefficients",
"anova",
"effects",
"significance",
"residual_diagnostics",
],
)
ms = result["model_summary"]
print(f"formula : {ms['formula']}")
print(f"R2 : {ms['r_squared']:.3f}")
print(f"R2 (adj) : {ms['r_squared_adj']:.3f}")
print(f"R2 (pred) : {ms['r_squared_pred']:.3f}")
print(f"obs / df_resid: {ms['n_obs']} / {ms['df_residual']}")
formula : y ~ (A + B + C + D) ** 2
R2 : 0.991
R2 (adj) : 0.979
R2 (pred) : 0.963
obs / df_resid: 19 / 8
Coefficient table#
Sort by absolute t-value to put the most impactful terms at the top. Coefficients with a small p-value are the ones that the data picks out as important; coefficients whose confidence interval straddles zero are not.
[5]:
coef_df = pd.DataFrame(result["coefficients"]).set_index("term")
coef_df = coef_df.reindex(coef_df["t_value"].abs().sort_values(ascending=False).index)
coef_df.round(3)
[5]:
| coefficient | std_error | t_value | p_value | ci_low | ci_high | |
|---|---|---|---|---|---|---|
| term | ||||||
| Intercept | 61.982 | 12.449 | 4.979 | 0.001 | 33.274 | 90.690 |
| B | -116.802 | 25.774 | -4.532 | 0.002 | -176.238 | -57.367 |
| A:B | 18.047 | 4.362 | 4.137 | 0.003 | 7.987 | 28.107 |
| A:D | -10.810 | 2.930 | -3.689 | 0.006 | -17.567 | -4.052 |
| D | 63.135 | 19.425 | 3.250 | 0.012 | 18.342 | 107.929 |
| A | -4.552 | 2.150 | -2.117 | 0.067 | -9.509 | 0.405 |
| C[T.Yes] | 9.418 | 4.552 | 2.069 | 0.072 | -1.079 | 19.914 |
| C[T.Yes]:D | 1.207 | 0.719 | 1.679 | 0.132 | -0.451 | 2.866 |
| B:D | 3.437 | 2.236 | 1.537 | 0.163 | -1.720 | 8.594 |
| B:C[T.Yes] | -1.741 | 1.151 | -1.512 | 0.169 | -4.395 | 0.913 |
| A:C[T.Yes] | -0.326 | 0.424 | -0.769 | 0.464 | -1.303 | 0.651 |
ANOVA#
[6]:
anova = pd.DataFrame(result["anova_table"]).set_index("source")
anova.round(3)
[6]:
| df | sum_sq | mean_sq | F | p_value | |
|---|---|---|---|---|---|
| source | |||||
| C | 1 | 217.794 | None | 651.088 | 0.000 |
| A | 1 | 16.460 | None | 49.206 | 0.000 |
| A:C | 1 | 0.198 | None | 0.591 | 0.464 |
| B | 1 | 2.502 | None | 7.479 | 0.026 |
| B:C | 1 | 0.765 | None | 2.287 | 0.169 |
| D | 1 | 9.319 | None | 27.858 | 0.001 |
| C:D | 1 | 0.943 | None | 2.818 | 0.132 |
| A:B | 1 | 5.725 | None | 17.114 | 0.003 |
| A:D | 1 | 4.552 | None | 13.608 | 0.006 |
| B:D | 1 | 0.790 | None | 2.362 | 0.163 |
| Residual | 8 | 2.676 | None | NaN | NaN |
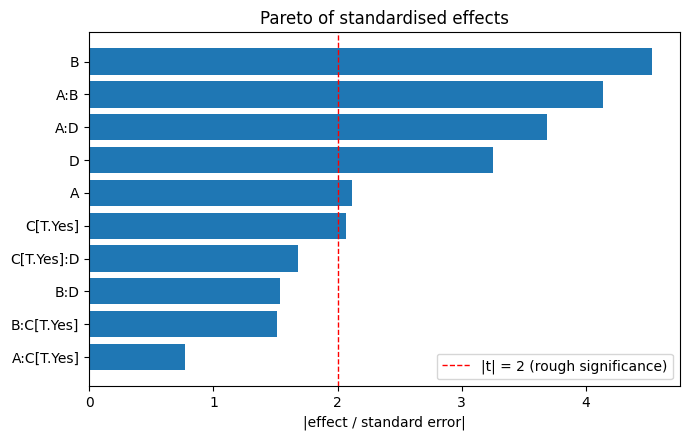
Pareto plot of standardised effects#
Dividing each effect estimate by its standard error gives a unitless comparison of how strong each term is relative to the noise. Plotting the absolute values as a horizontal bar chart from largest to smallest is the classical Pareto plot.
[7]:
effects = pd.Series(result["effects"], dtype=float)
errors = pd.Series(result["effect_std_errors"], dtype=float).reindex(effects.index)
standardised = (effects / errors).abs().sort_values()
_fig, ax = plt.subplots(figsize=(7, 0.35 * max(4, len(standardised)) + 1.0))
ax.barh(standardised.index, standardised.values, color="#1f77b4")
ax.axvline(2, color="r", linestyle="--", linewidth=1, label="|t| = 2 (rough significance)")
ax.set_xlabel("|effect / standard error|")
ax.set_title("Pareto of standardised effects")
ax.legend(loc="best")
plt.tight_layout()
plt.show()
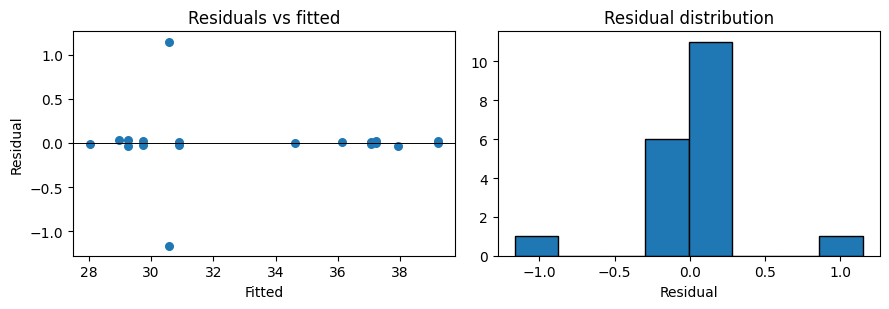
Residual diagnostics#
[8]:
diag = result["residual_diagnostics"]
fitted = np.asarray(diag["fitted_values"], dtype=float)
resid = np.asarray(diag["residuals"], dtype=float)
_fig, axes = plt.subplots(1, 2, figsize=(9, 3.2))
axes[0].scatter(fitted, resid, s=30, color="#1f77b4")
axes[0].axhline(0, color="k", linewidth=0.7)
axes[0].set_xlabel("Fitted")
axes[0].set_ylabel("Residual")
axes[0].set_title("Residuals vs fitted")
axes[1].hist(resid, bins=8, color="#1f77b4", edgecolor="k")
axes[1].set_xlabel("Residual")
axes[1].set_title("Residual distribution")
plt.tight_layout()
plt.show()
print(f"Shapiro-Wilk p-value : {diag['shapiro_wilk']['p_value']:.3f}")
print(f"Durbin-Watson : {diag['durbin_watson']:.2f}")
print(f"Breusch-Pagan p-value: {diag['breusch_pagan']['p_value']:.3f}")
Shapiro-Wilk p-value : 0.000
Durbin-Watson : 1.52
Breusch-Pagan p-value: 0.050
What to try next#
Drop the terms below the
|t| = 2line and re-fit. The model R squared will go down a touch but the adjusted R squared usually goes up because the noise terms penalise the unadjusted statistic.Pass
analysis_type="lack_of_fit"to ask whether the residual variation is consistent with pure replication error, which tells you whether a curvature term is missing.For follow-up experimentation, the :func:
process_improve.experiments.strategy.recommend_strategyfunction will propose where to run the next batch of experiments given the coefficients you have already estimated.